Introduction
The human genome sequence was completed in draft form in 2001 (Lander et al., 2001; Venter et al., 2001). Shortly thereafter, the genome sequences of several model organisms were determined (Chinwalla et al., 2002; Gibbs et al., 2004; The Chimpanzee Sequencing and Analysis Consortium, 2005). These feats were accomplished with Sanger DNA sequencing, which was limited in throughput and high cost; indeed the first human genome sequence was estimated to cost 0.5–1 billion dollars. These limitations reduced the potential of DNA sequencing for other applications, such as personal genome sequencing. Following the release of the “finished” human genome (International Human Genome Sequencing Consortium, 2004), the National Human Genome Research Institute (NGHRI) created a 70 million dollar DNA sequencing technology initiative aimed at achieving a $1000 human genome in ten years (Schloss, 2008), and a flurry of high-throughput sequencing (HTS) technologies emerged.
To put this initiative in perspective, improvements to traditional Sanger sequencing had decreased the per base cost by around 100-fold by the completion of the Human Genome Project (Schloss, 2008). To reach the $1000 dollar genome threshold, however, an additional leap of 5 orders of magnitude was necessary. Much of this divide has been traversed—the cost of a genome sequence (without interpretation) is presently less than $2,000. The road to this milestone involved many commercial HTS platforms, which differ in their details but typically follow a similar general paradigm: template preparation, clonal amplification, followed by cyclical rounds of massively parallel sequencing. The specific strategy employed by each platform determines the quality, quantity and biases of the resulting sequence data and the platform’s usefulness for particular applications.
Several excellent reviews have covered HTS platform strategies in great depth (Metzker, 2010; Morey et al., 2013). Many important platforms are not covered here, including Roche/454’s pyrophosphate Genome Sequencer (Margulies, 2005), Helicos’ single molecule Heliscope sequencer (Harris et al., 2008) as well as the Polonator (Shendure et al., 2005), ABI’s SOLiD (Valouev et al., 2008) and Complete Genomics’ DNA nano-array sequencer (Drmanac et al., 2010). Instead, we focus on the most commonly used platforms today as well as more recent developments. We also provide an overview of the growing array of HTS applications and highlight their use by the genomics community to illuminate previously intractable topics in biology. Finally, we discuss the limitations of current platforms and challenges to clinical sequencing.
Overview of selected commercially available high-throughput sequencing platforms
Illumina
Illumina/Solexa released the Genome Analyzer II in 2006, and advances in Illumina’s technology over the intervening years have largely set the pace for the tremendous gains in output and reductions in cost (Figure 1). As a consequence, Illumina machines currently dominate the HTS market. The sequencing process involves clonal amplification of adaptor-ligated DNA fragments on the surface of a glass slide (Bentley et al., 2008) (Figure 2A). Bases are read using a cyclic reversible termination strategy, which sequences the template strand one nucleotide at a time through progressive rounds of base incorporation, washing, imaging and cleavage. In this strategy, fluorescently-labeled 3´-O-azidomethyldNTPs are used to pause the polymerization reaction, enabling removal of unincorporated bases and fluorescent imaging to determine the added nucleotide (Guo et al., 2008). Following scanning of the flow cell with a coupled-charge device (CCD) camera, the fluorescent moiety and the 3´ block are removed, and the process is repeated. Across all Illumina models, the overall error rates are below 1%, and the most common type of error is substitutions (Dohm et al., 2008).
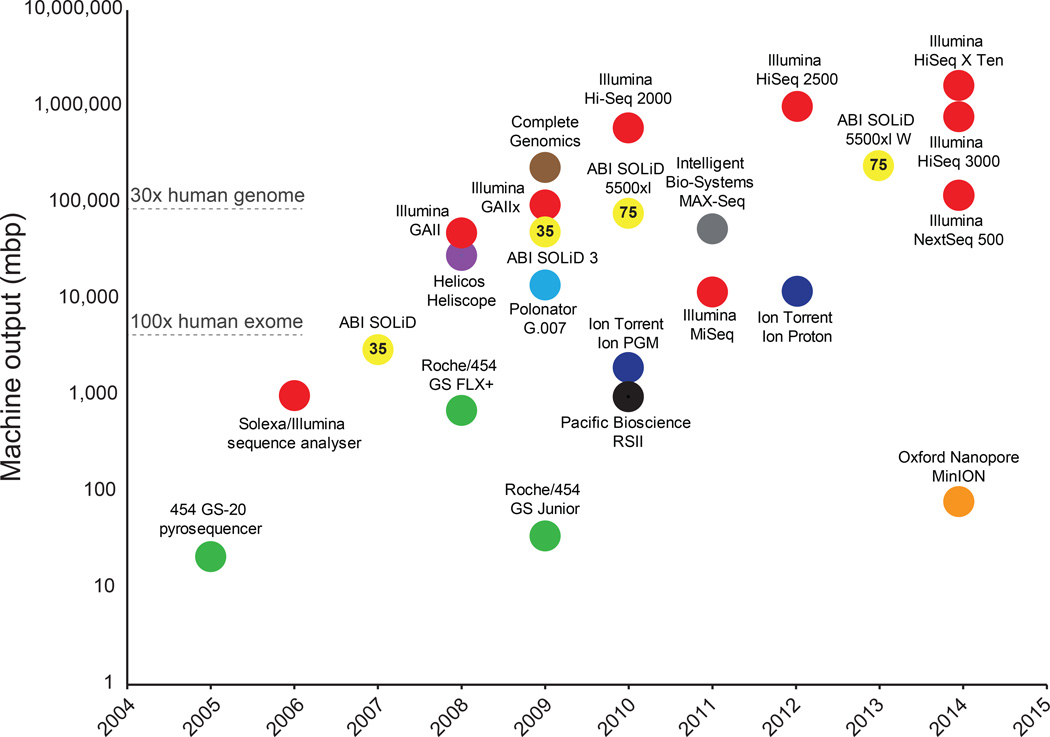
Figure 1. Timeline and comparison of commercial HTS instruments.
Plot of commercial release dates versus machine outputs per run are shown. For the MinION, outputs from an 18 hour run were used (Ashton et al., 2014). Numbers inside data points denote current read lengths. Sequencing platforms are color-coded.
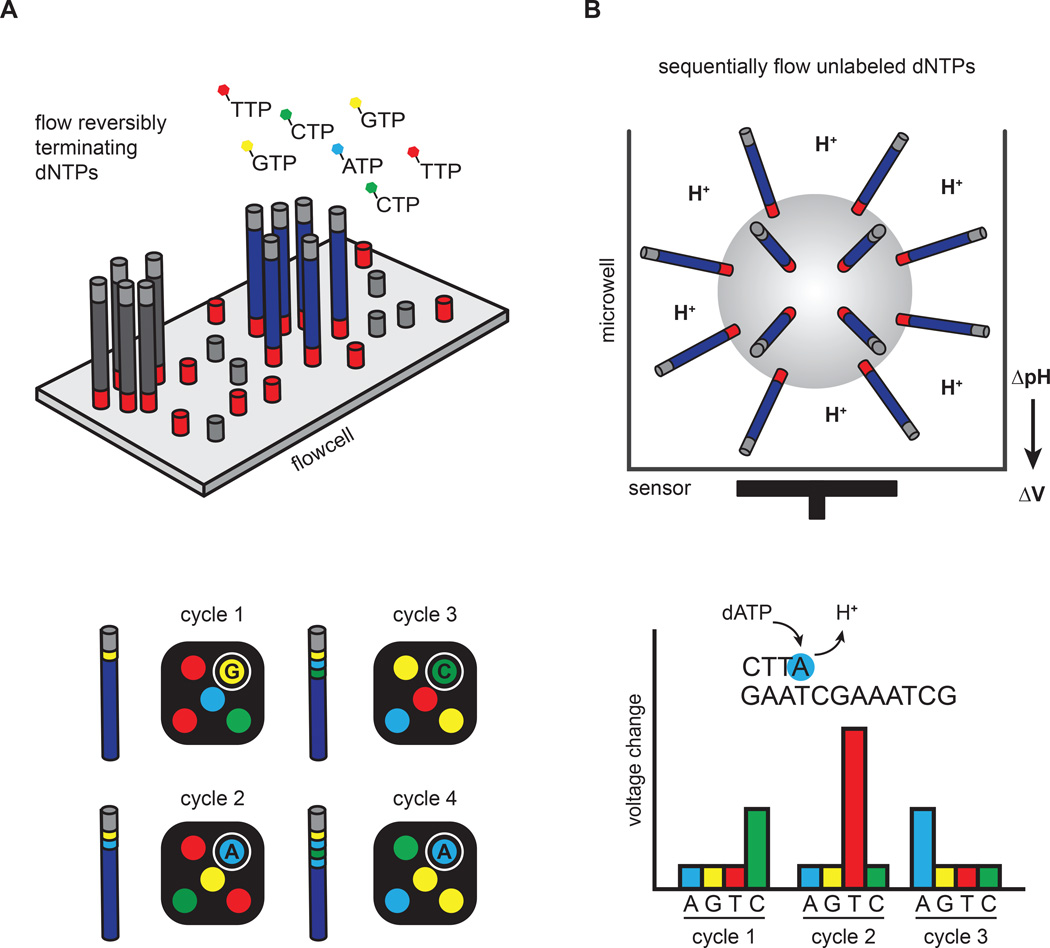
Figure 2. Clonal amplification-based sequencing platforms.
(A) Illumina’s four-color reversible termination sequencing method. DNA templates are first clonally amplified on the surface of a glass flow cell. Sequencing occurs via successive rounds of base incorporation, washing and imaging. A cleavage step after image acquisition removes the fluorescent dye and regenerates the 3´OH for the next cycle. Analysis of four-color images is used to determine base composition. (B) Ion Torrent’s semiconductor sequencing method. Emulsion-PCR is used to clonally amplify DNA templates on the surface of beads, which are subsequently placed into microwells. pH changes induced by the release of hydrogen ions during DNA extension are detected by a sensor positioned at the bottom of the microwell. These pH changes are converted into a voltage signal, which is proportional to the number of nucleotides added by the polymerase.
Illumina currently produces a suite of sequencers (MiSeq, NextSeq 500 and the HiSeq series) optimized for a variety of throughputs and turnaround times. The MiSeq and HiSeqs are the most established platforms. The MiSeq is designed as a fast, personal benchtop sequencer, with run times as low as 4 hours and outputs intended for targeted sequencing and sequencing of small genomes. The HiSeq 2500, on the other hand, is engineered for high-throughput applications, yielding current outputs of 1 Tb in 6 days. Unlike previous HiSeq models, the HiSeq 2500 can also be run in rapid mode, which is less cost effective but can produce a 30x human genome in 27 hours.
In early 2014, Illumina introduced the NextSeq 500 as well as the HiSeq X Ten. Similar to the MiSeq, the NextSeq 500 is designed as a fast benchtop sequencer for individual labs. However, the NextSeq is capable producing 120 Gb, or a single 30x genome, in less than 30 hours. The NextSeq 500 system also employs a novel two-channel sequencing strategy. In this approach, cytosine is labeled red, thymine is labeled green, adenine is effectively yellow (labeled with a mixture of red and green) and guanine is unlabeled. In contrast to the four-channel strategy used in the MiSeq and HiSeq platforms, two-channel sequencing requires only two images for nucleotide detection, reducing data processing times and increasing throughput. Despite the reduced complexity, the overall error rates (<1%) are similar to the more established HiSeq machines.
The HiSeq X Ten is a population-scale whole genome sequencing (WGS) system that was also released in 2014. It is capable of outputting 1.8T in 3 days or 18,000 genomes at 30x coverage per year. Currently, Illumina only supports WGS of human samples on HiSeq X Ten systems. In addition to enhanced optics and computing capacity, the HiSeq X Ten dramatically increases throughput by incorporating a new patterned flow cell technology that improves cluster generation chemistry. Patterned flow cells contain billions of nanowells that standardize cluster spacing and size, allowing higher cluster densities. Patterned flow cell technology is also used in the recently released HiSeq 3000/4000 machines, which provide outputs and run times in between the HiSeq X Ten and the HiSeq 2500.
Life Technologies/ThermoFisher/Ion Torrent
Life Technologies commercialized Ion Torrent’s semiconductor sequencing technology in 2010 in the form of the benchtop Ion PGM sequencer. The template preparation and sequencing steps are conceptually similar to the Roche/454 pyrosequencing platform (Margulies, 2005). Namely, emulsion-PCR is used to clonally amplify adapter-ligated DNA fragments on the surface of beads. The beads are subsequently distributed into microwells where a sequencing-by-synthesis reaction occurs (Figure 2B). Unlike pyrosequencing, which couples base incorporation with luciferase-based light production, Ion Torrent’s semiconductor sequencing measures pH changes induced by the release of hydrogen ions during DNA extension (Rothberg et al., 2011). These pH changes are detected by a sensor positioned at the bottom of the microwell and converted into a voltage signal. The voltage signal is proportional to the number of bases incorporated, and the sequential addition of individual nucleotides during each sequencing cycle allows base discrimination. Moreover, Ion Torrent avoids optical scanning to distinguish nucleotides during cycles of sequencing, a difference that dramatically speeds sequencing runs and reduces costs.
Ion Torrent released a second machine in 2012, the Ion Proton, which increases output over the PGM by an order of magnitude (1Gb versus 10Gb). However, the Proton currently features a maximum of 200bp read lengths as opposed to 400bp for the PGM. Multiple chips are also available to tailor outputs for different applications. The PGM is most useful for targeted resequencing projects and small genome analysis, whereas the Proton is capable of exome sequencing and whole-transcriptome analysis. The speed of sequencing, 2–8 hours depending on the machine and chip used, make these sequencers particularly useful for clinical applications (Mellmann et al., 2011). Insertions and deletions (indels) are the most common error types (Liu et al., 2012). Because the correlation between the number of bases incorporated and the subsequent voltage change does not perfectly scale, homopolymer repeats longer than 6 base pairs lead to increased error rates (Rothberg et al., 2011).
Pacific Biosciences
Single-molecule real-time (SMRT) sequencing was pioneered by Nanofluidics, Inc. and commercialized by Pacific Biosciences. Template preparation involves ligation of single-stranded, hairpin adapters onto the ends of digested DNA or cDNA molecules, generating a capped template (SMRT-bell). By using a strand displacing polymerase, the original DNA molecule can be sequenced multiple times, thereby increasing accuracy (Travers et al., 2010). Importantly, clonal amplification is avoided, allowing direct sequencing of native, and potentially modified, DNA. DNA synthesis occurs in zeptoliter-sized chambers, called zero-mode waveguides (ZMW), in which a single polymerase is immobilized at the bottom of the chamber (Levene et al., 2003) (Figure 3A). The physics of these chambers reduces background noise such that phosphate-labeled versions of all 4 nucleotides can be present simultaneously. Thus, polymerization occurs continuously, and the DNA sequence can be read in real-time from the fluorescent signals recorded in a video (Eid et al., 2009).
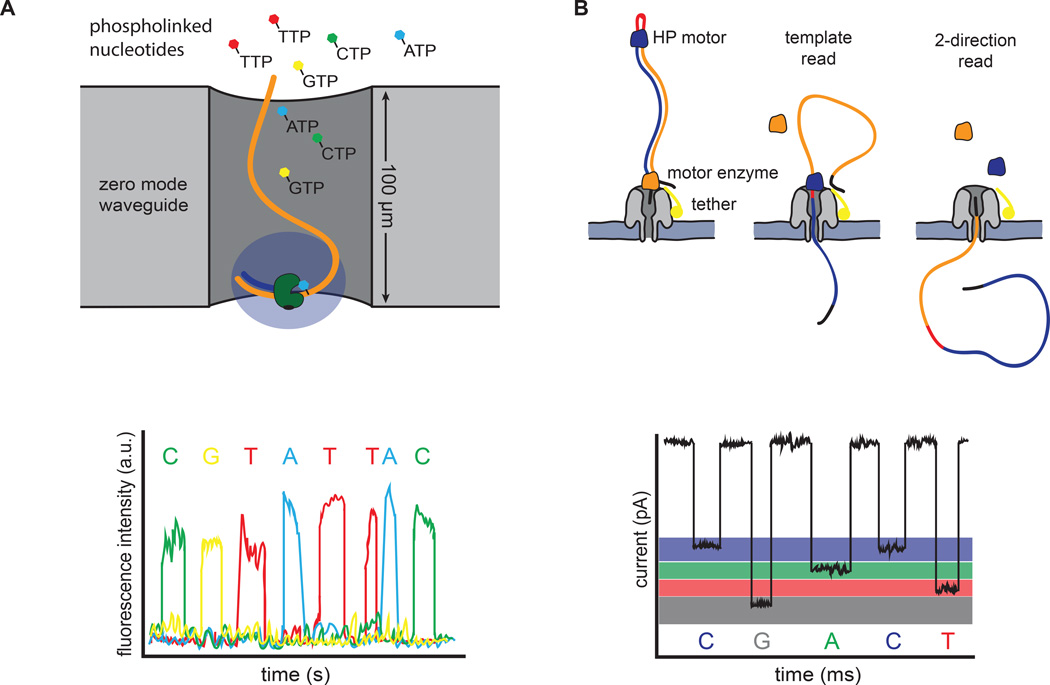
Figure 3. Single molecule sequencing platforms.
(A) Pacific Bioscience’s SMRT sequencing. A single polymerase is positioned at the bottom of a zero-mode waveguide (ZMW). Phosphate-labeled versions of all 4 nucleotides are present, allowing continuous polymerization of a DNA template. Base incorporation increases the residence time of the nucleotide in the ZMW, resulting in a detectable fluorescent signal that is captured in a video. (B) Oxford Nanopore’s sequencing strategy. DNA templates are ligated with two adapters. The first adapter is bound with a motor enzyme as well as a tether, whereas the second adapter is a hairpin oligo that is bound by the HP motor protein. Changes in current that are induced as the nucleotides pass through the pore are used to discriminate bases. The library design allows sequencing of both strands of DNA from a single molecule (2-direction reads).
Released in 2010, the RS II remains Pacific Biosciences only commercially available machine. However, altering the chemistry and doubling the number of ZMWs to 150k per SMRT cell have greatly enhanced performance. Using the latest chemistry, each SMRT cell produces ~50k reads and up to 1Gb of data in 4 hours. The average read lengths are >14kb, but individual reads can be as long 60kb. As with most single molecule sequencing platforms, high error rates (~11%) are evident for single pass reads, and these errors are dominated by indels. Sequencing errors, however, are distributed randomly, allowing accurate consensus calls with increasing coverage or multiple passes around the same template, socalled circular consensus sequences (Carneiro et al., 2012; Koren et al., 2012). By avoiding clonal amplification, SMRT sequencing is also much less sensitive to GC sequence content than other platforms (Loomis et al., 2013). This suite of characteristics makes SMRT sequencing particularly useful for projects involving de novo assembly of small bacterial and viral genomes as well as large genome finishing (English et al., 2012). Reconstructing structural variation in the genome (Chaisson et al., 2014) and isoform usage in the transcriptome (Sharon et al., 2013) are also key areas where SMRT sequencing has clear advantages over short read technologies. However, lower throughput and higher per base sequencing costs currently limit the scope of most genome-wide studies.
In addition to providing long, unbiased reads, another distinguishing characteristic of SMRT sequencing is that the polymerization reaction is monitored in real-time, allowing data pertaining to both base composition and enzymatic kinetics to be collected. Distinct kinetic profiles are produced as the polymerase encounters various types of DNA methylation (Flusberg et al., 2010). These kinetic signatures have been utilized to map sites of potential 6-methyladenine and 5-methylcytosine genomewide in bacteria (Fang et al., 2012). It is possible that these approaches will be extended to map other types of DNA modifications, including DNA damage induced in cancer cells. Moreover, SMRT sequencing instruments are not limited to studying DNA alone, as other molecules, such as ribosomes, can be tethered to the bottom of the ZMW and monitored at single molecule resolution (Uemura et al., 2010).
Oxford Nanopore Technologies
Nanopore-based sequencing is an emerging single molecule strategy that has made significant progress in recent years, with Oxford Nanopore Technologies leading the development and commercialization of this method. Nanopore sequencing can take a variety of forms, but principally relies on the transition of DNA or individual nucleotides through a small channel (Wang et al., 2015). In Oxford Nanopore’s current technology, a sequencing flow cell comprises hundreds of independent micro-wells, each containing a synthetic bilayer perforated by biologic nanopores. Sequencing is accomplished by measuring characteristic changes in current that are induced as the bases are threaded through the pore by a molecular motor protein (Figure 3B). Library preparation is minimal, involving fragmentation of DNA and ligation of adapters. Much like SMRT sequencing, this library preparation methodology can be done with or without PCR-amplification. The first adapter is bound with a propriety motor enzyme as well as a molecular tether, whereas the second adapter is a hairpin oligonucleotide that is bound by a second socalled HP motor protein (Quick et al., 2014). This library design allows sequencing of both strands of DNA from a single molecule, which increases accuracy (Ashton et al., 2014; Quick et al., 2014).
The first commercially available device for nanopore sequencing is the MinION, a USB-powered, portable sequencer, which Oxford Nanopore Technologies released in early 2014 as part of an early access program. A single 18 hour run can produce >90 Mbp of data from around 16,000 total reads, with median and maximum read lengths of ~6 kb and >60 kb, respectively (Ashton et al., 2014). As with all single molecule sequencing methodologies, error rates are high. Jain and colleagues most recently reported insertion, deletion and substitution rates of 4.9%, 7.8% and 5.1%, respectively (Jain et al., 2015). Presently, it also has a very high run failure rate. Despite the high error rates, MinION reads have been successfully used to determine the position and structure of a bacterial resistance island in combination with Illumina-derived reads (Ashton et al., 2014) and resolve an assembly gap on human Xq24 (Jain et al., 2015). Given the relatively high error rates and low throughput, nanopore sequencing is unlikely to overtake current sequencing platforms in the near future; however, the combination of size, speed, read lengths and machine cost hold promise for the future.
The development and use of HTS applications
As sequencing costs have fallen, HTS machines have become widely present in university core facilities and even individual labs. Decreasing costs and increased accessibility have enabled researchers to develop a rich catalog of HTS applications (Figure 4 and Table 1). Some of these technologies were initially developed using DNA microarrays, but many are enabled only by using sequencing. High-throughput sequencing offers many advantages over DNA microarrays. In particular, it is more precise and not subject to cross-hybridization, thereby providing higher accuracy and a larger dynamic range (>105 for DNA sequencing vs 102 for DNA microarrays) (Wang et al., 2009). Similar to microarrays, however, HTS-based applications can be biased by a number of variables, such as sequencing platform and library preparation method. The Sequencing Quality Control Consortium and similar initiatives are designed to study these biases and develop approaches to control for them, as has been recently demonstrated for RNA-seq (Su et al., 2014).
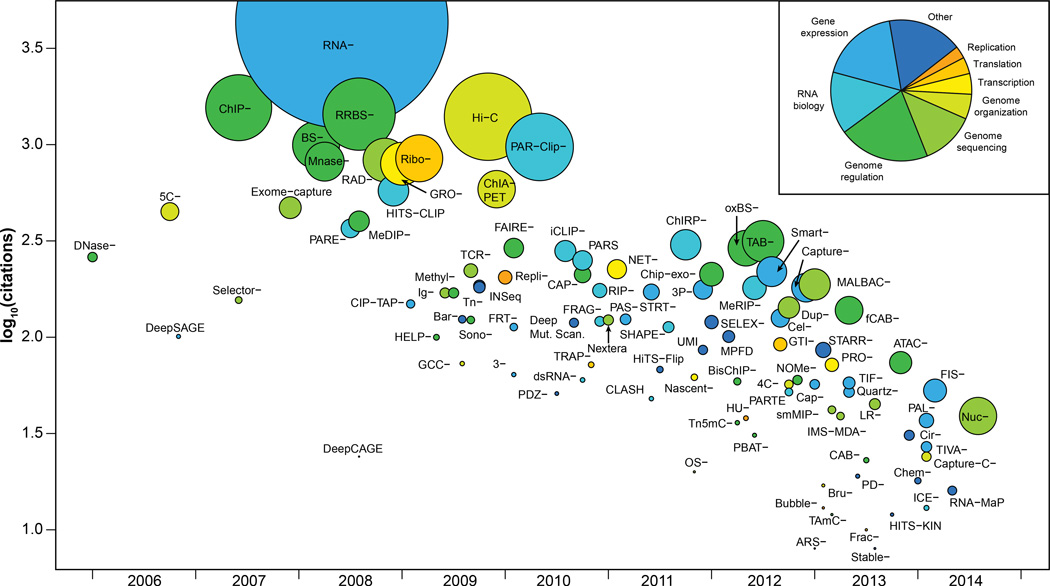
Figure 4. Broad overview of HTS applications.
Publication date of a founding article describing a method versus the number of citations that the article received. Methods are colored by category, and the size of the data point is proportional to publication rate (citations/months). The inset indicates the color key as well the proportion of methods in each group. For clarity, Seq has omitted from the labels.
Table 1.
| Method | Purpose | Reference |
|---|---|---|
| RNA-Seq | Transcript analysis | (Nagalakshmi, U., et al., 2008) |
| Global run-on sequencing (GRO-Seq) | Transcription | (Core, L. J., et al., 2008) |
| Nascent-Seq | Transcription | (Khodor, Y.L., et al., 2011) |
| Native elongating transcript sequencing (NET-Seq) | Transcription | (Churchman, L., et al., 2011) |
| Ribo-Seq | Translation | (Ingolia, N.T., et al., 2009) |
| Replication sequencing (Repli-Seq) | Replication | (Hansen, R.S., et al., 2010) |
| Hi-C | Chromatin conformation | (Lieberman-Aiden, E., et al., 2009) |
| Chromatin interaction analysis by paired-end tag sequencing (ChIA-PET) | Chromatin conformation | (Fullwood, M.J., et al., 2009) |
| 5-C-Seq | Chromatin conformation | (Dotsie, J., et al., 2006) |
| Chromatin isolation by RNA purification sequencing (ChIRP-Seq) | Genome localization | (Chu, C., et al., 2011) |
| Reduced representation bisulphite sequencing (RRBS-Seq) | Genome methylation | (Meissner, A., et al., 2008) |
| Bisulfite sequencing (BS-Seq) | Genome methylation | (Cokus, S.J., et al., 2008) |
| DNAse-Seq | Open chromatin | (Crawford, G.E., et al., 2006) |
| Assay for transposase-accessible chromatin using sequencing (ATAC-Seq) | Open chromatin | (Buenrostro, J.D., et al., 2013) |
| Parallel Analysis of RNA structure (PARS) | RNA structure | (Wan, Y., et al., 2012) |
| Structure-Seq | RNA structure | (Ding, Y., et al., 2014) |
| RNA on a massively parallel array (RNA-MaP) | RNA-protein interactions | (Buenrostro, J.D., et al., 2014) |
| RNA immunoprecipitation sequencing (RIP-Seq) | RNA-protein interactions | (Sephton, C.F., et al., 2010) |
| Parallel analysis of RNA ends sequencing (PARE-Seq) | microRNA target discovery | (German, M.A., et al., 2008) |
| Massively parallel functional dissection sequencing (MPFD) | Enhancer assay | (Patwardhan, R.P., et al., 2012) |
As HTS-based applications have become more robust, they have not only enabled individual researchers but also a variety of consortia-based projects. These large-scale projects have both provided valuable resources to the community and also have addressed questions that would be difficult for individual labs to approach. Some of these projects are listed in Table 2 and include efforts to characterize the human genome (ENCODE, Roadmap Epigenomics Project), study human genetic variation (1000 Genomes Project), analyze gene expression (GTEx), and discover the molecular underpinnings of human disease (many; see Table 2). These coordinated efforts produce foundational resources that are of high utility to the scientific community by depositing the data into easily accessible public databases. Moreover, consortia often implement robust experimental and computational standards (e.g., Landt et al., 2012), ensuring high quality data. Use of HTS applications by both individual laboratories and the large consortia have enabled researchers to illuminate previously intractable topics in biology, some of which are discussed below.
Table 2.
Examples of consortia-based projects
| Initiative | Purpose | Website |
|---|---|---|
| 1000 Genomes Project | Cataloging normal variation in diverse human populations. | www.1000genomes.org |
| The Encyclopedia of DNA Elements | Identifying functional genomic elements in the human genome. | www.encodeproject.org |
| Roadmap Epigenomics Project | Catalogue human epigenomic data with the goal of advancing basic biology and disease-oriented research. | www.roadmapepigenomics.org |
| Human Microbiome Project | Comprehensive characterization of the human microbiome and analysis of its role in human health and disease. | www.hmpdacc.org |
| Genotype-Tissue Expression Program | Characterizing gene expression and regulation in many human tissues and correlating with genetic variation and disease. | www.commonfund.nih.gov/GTEx/index |
| Human Immunology Project Consortium | Characterizing the diverse states of the human immune system following infection, vaccination or treatment. | http://www.immuneprofiling.org |
| Grand Opportunity Exome Sequencing Project | Discovery of novel genes and mechanisms contributing to heart, lung and blood disorders. | https://esp.gs.washington.edu/drupal |
| The Cancer Genome Atlas | Understanding the molecular basis of cancer. | www.cancergenome.nih.gov |
| International Cancer Genome Consortium | Describing the genomic, transcriptomic and epigenomic changes in 50 different tumor types. | www.icgc.org |
| Clinical Sequencing Exploratory Research Program | Develop methods as well as the legal and ethical frameworks necessary to integrate sequencing into the clinic. | www.genome.gov/27546194 |
| Centers for Mendelian Genomics | Discovering the genes and genetic variants underlying human Mendelian disorders. | www.mendelian.org |
| Undiagnosed Diseases Network | Promoting the use of genomic data to elucidate the mechanisms underlying the diseases of unknown etiology. | www.commonfund.nih.gov/Diseases/index |
| Newborn Sequencing in Genomic Medicine and Public Health | Exploring the challenges and opportunities associated with using genomic sequence information in the newborn period. | www.genome.gov/27558493 |
| The Pediatric Cardiac Genomics Consortium | Determining the genes responsible for congenital heart disease. | www.benchtobassinet.com |
| Alzheimer's Disease Sequencing Project | Identifying genes contributing to risk of developing Alzheimer's disease in multiethnic populations. | www.niagads.org/adsp |
Genome sequencing and variation
The utility of HTS technologies for determining genome sequences de novo was first demonstrated by sequencing the genome of Acinetobacter baumannii (Smith et al., 2007). As the technologies and throughput improved, they were applied to "resequencing" human genomes and exomes, which was accomplished by first mapping reads to a reference genome and then identifying variants that differ between the sample genome and the reference (Wheeler et al., 2008). The different genome sequencing projects have since revealed that individuals typically harbor 3.5–4 million single nucleotide variants (SNVs) in total and several hundred thousand short indels relative to the reference genome. Importantly, these variants include hundreds of loss of function alterations in genes (The 1000 Genomes Project Consortium, 2010).
HTS has also been used to globally characterize structural variation (SV) in the human genome. SVs include large (>1kb) segments of the genome that have been duplicated, deleted or rearranged. The short read lengths of most HTS platforms make determining SVs and indels more challenging than SNVs (Snyder et al., 2010). Typically, at least four independent approaches are utilized to identify SVs in a genome. These approaches include depth of read coverage (Abyzov et al., 2011), mapping of paired end reads that are discordant from the reference genome (Korbel et al., 2007), identifying split reads (Zhang et al., 2011) and mapping of breakpoint junctions (Kidd et al., 2010). Although each method has shortcomings, the improvement in resolution over array-based approaches has greatly enhanced our understanding of the prevalence of SVs throughout the genome and their contribution to disease. However, because no method or combination of them is comprehensive, SVs are never characterized in their entirety, if at all, in most sequencing projects.
In addition to identifying variants, it is also useful to assign them to paternal and maternal alleles, or “phase” them. Similar to SVs, current read lengths hinder our ability to phase genomes. This limitation can be circumvented by several methods, including sequencing parents, sequencing proximity ligated fragments (Selvaraj et al., 2013) or dilution and barcoding strategies during template preparation to allow long read assembly (Kuleshov et al., 2014; Voskoboynik et al., 2013). With approximately 30 Gbp of additional sequence data, ~99% of the SNVs identified in a 50x genome can be phased into blocks that are 0.2–1Mb in length (Kuleshov et al., 2014). Understanding the phase of variants can have important clinical implications when determining if multiple damaging variants affect both copies of a gene or only one copy. To date, HTS has been applied to many thousands of genomes and many tens of thousands of exomes, yielding tremendous insight into human diversity and disease.
Mapping regulatory information of the genome
HTS has applications beyond simply sequencing genomes. Perhaps one of the highest impact areas is the genome-wide mapping of DNA regulatory elements at high-resolution. The first of these technologies was ChIP-Seq in which DNA associated with a transcription factor (TF) or chromatin modification is immuno-selected and then sequenced using HTS (Johnson et al., 2007). Mapping the sequences back to the genome reveals the location of bound regions or chromatin modifications. A more general method for discovering many putative regulatory regions is to map “open” regions of the genome using DNase I digestion, followed by DNA sequencing of the ends of the fragments (Crawford et al., 2006). This method identifies approximately 50% of regions that are TF-bound as measured by ChIP-Seq (Cheng et al., 2014). DNase-Seq, however, is quickly being replaced by Assay for Transposon Accessible Chromatin-Seq (ATAC-Seq) in which transposon-based insertion is used to map open chromatin regions with approximately 50 million mapped reads (Buenrostro et al., 2013). The ATAC-seq protocol is also simpler and can be applied to small numbers of cells, even single cells.
Regulatory information is especially revealing when compared across many individual genomes or within a single genome across many cell or tissue types. Large-scale application of these methods by the ENCODE project has provided a wealth of invaluable information regarding transcription factor binding networks (The ENCODE Project Consortium, 2012), epigenetic maps (Thurman et al., 2012) and transcript annotations (Djebali et al., 2012). Moreover, recent studies have found more than 3.5 million regulatory elements located throughout the genome in different cell types (Roadmap Epigenomics Consortium et al., 2015). One of the most striking findings from these studies as a whole, however, was the higher than expected portion of the genome that appears to be functional. The exact percentage is a source of significant debate (Doolittle, 2013), highlighting the importance of further experimental evidence to assign function to genomic elements. Genome targeting techniques, such as CRISPR-Cas9 (Gilbert et al., 2014) as well as high-throughput enhancer assays (Kheradpour et al., 2013) provide researchers with new tools to interrogate putative regulatory elements. Nonetheless, a variety of lines of evidence (GWAS, ENCODE) suggest that the total amount of regulatory regions is likely greater than that of protein coding regions (Kellis et al., 2014).
Mapping the three-dimensional organization of the genome
Our understanding of the global organization and compartmentalization of chromosomes has been profoundly advanced by HTS technologies. 3D chromatin interactions can be studied using a variety of HTS assays, such as ChIA-PET (chromatin interaction analysis by paired-end tag sequencing) and Hi-C (Fullwood et al., 2009; Lieberman-Aiden et al., 2009). Each of these assays relies upon proximity-based ligation of cross-linked, sheared chromatin followed by sequencing to derive contact maps. Hi-C was the first technique to allow unbiased, genome-wide interrogation of chromatin organization and revealed that the genome broadly partitions into open and closed chromatin states (Lieberman-Aiden et al., 2009). Hi-C also demonstrated that the genome is organized into topological associating domains (TADs), which show high amounts of intra-domain interactions but exhibit infrequent interactions across domain boundaries (Dixon et al., 2012). Interestingly, TAD organization is stable across cell types and evolutionarily conserved across species. The boundaries between TADs were also enriched for housekeeping genes and binding sites for the insulator protein CTCF, raising the possibility that the distribution of TADs is chromosomally encoded (Dixon et al., 2012).
Recent advancements to the Hi-C technique combined with extremely deep sequencing (billions of reads per sample) have produced much higher resolution contact maps (~1kb), which refine TAD domain size from 1 Mb to less than 200 kbp (Rao et al., 2014). These new Hi-C maps demonstrated intrachromosomal looping events, often containing promoter-to-enhancer contacts that were associated with gene activation. Most loops were anchored with directionally-oriented CTCF binding sites, suggesting a mechanistic role for CTCF in establishing stable loops. Strikingly, fewer than 10,000 looping events were observed genome-wide, which is far smaller than previous estimates (Jin et al., 2013). Modeling of Hi-C data has also suggested a fractal globule chromatin state, a conformation that both maximizes packing while preserving the flexibility to access any genomic locus (Lieberman-Aiden et al., 2009).
Characterizing the transcriptome
Our appreciation for the diverse cellular roles of RNA has been greatly enhanced by the advent of high-throughput sequencing. Much of this evolution in thought has been a direct result of the many HTS applications designed to systematically identify various classes of RNA as well as to characterize RNA structure, RNA-protein interactions and genomic localization. Cap analysis of gene expression (CAGE) and RNA-seq have been utilized to great effect to deeply characterize transcriptomes, providing precise, comprehensive measurements for message abundance, isoform usage, RNA-editing and allele-specific expression. Deep sequencing of RNA has suggested that roughly three-quarters of the human genome is transcribed (Djebali et al., 2012). Most of this transcription covers introns or is very low, non-coding and of unclear biologic significance. However, many interesting species of non-coding RNA, including lncRNAs (long, non-coding), snoRNAs (small, nucleolar) and microRNAs, have been systematically described with RNA-seq and derivative techniques. A subset of lncRNAs, for example, have been revealed by overlaying RNA-seq data with ChIP-seq profiles characteristic of expressed genes (Guttman et al., 2009). Building upon earlier cDNA sequencing and tiling array experiments, these HTS approaches expanded the list to include more than a thousand mammalian lncRNAs. Analogous expansions have occurred for many aspects of RNA biology, such as the number of sites undergoing RNA editing (Li et al., 2009).
Understanding the structure and biology of these newly discovered transcripts has led to the development of additional HTS applications. For instance, microRNA-target discovery has been facilitated by sequencing signatures of miRNA-mediated mRNA decay, using parallel analysis of RNA ends (PARE) (German et al., 2008). Furthermore, RNA immunoprecipitation chip (RIP-chip) and subsequently RIP-seq were utilized to show that approximately 20% of the lncRNAs associate with polycomb repressor complex 2 (PRC2), a chromatin-modifying complex (Khalil et al., 2009; Zhao et al., 2010). Given these links to chromatin, methods analogous to ChIP-seq were developed, such as chromatin isolation by RNA purification (ChIRP-seq), to determine the genomic localization of lncRNAs (Chu et al., 2011). HTS applications have also made it possible to determine transcript structure both in vitro (parallel analysis of RNA sequencing; PARS) and in vivo (Structure-seq), providing insight into the effects of various structural features on translation efficiency, splicing and polyadenylation (Ding et al., 2014; Kertesz et al., 2010). More recently, systematic interrogation of sequence-function relationships for RNA-protein interactions has been made possible using a high-throughput biochemical assay called RNA on a massively parallel array (RNA-MaP) (Buenrostro et al., 2014). The use of these assays, and many others, have enabled researchers to study RNA biology both comprehensively and with great detail, thereby enhancing our appreciation for the varied roles RNA plays in normal cellular homeostasis as well as human disease.
Microbiome sequencing
Advances in HTS have enabled extensive cataloging of metagenomic samples, providing insight into the diversity of microbial species from a wide variety of sources, including the ocean, soil and the human body. These studies use both 16S rRNA gene sequencing to determine phylogenetic relationships as well as more comprehensive shotgun sequencing to predict detailed species and gene composition. In particular, much attention has been paid to characterizing the diverse microbes resident to healthy human populations (The Human Microbiome Project Consortium, 2012). These studies found extensive variation in both body site habitat and among different individuals, giving rise to the concept of a “personal microbiome”. Microbial diversity, or the number and abundance distribution of microorganisms in a given niche, also correlates with several human diseases. For instance, an increase in diversity is associated with bacterial vaginosis (Fredricks et al., 2005), whereas obesity and inflammatory bowel disease exhibit a decrease in the diversity of gut microbes (Qin et al., 2010; Turnbaugh et al., 2009). Although transplant studies in mice have demonstrated a direct link between the gut microbiome, energy metabolism and obesity (Turnbaugh et al., 2006), causal relationships for the majority of human diseases are not well-established. A deeper understanding will require more detailed characterizations of the dynamics of microbiomes across health states as well as more integrative studies to investigate the functional interplay between the microbiota, the host and the environment.
Genome sequencing of rare diseases
The capacity to sequence genomes, exomes and transcriptomes has profoundly influenced our understanding of the genetics of human disease, especially for rare Mendelian disorders and cancer. According to the Online Mendelian Inheritance in Man database, there are more than 7800 Mendelian disorders, but the causative gene for less than one half of these are known. By sequencing unrelated patients or affected and unaffected family members, early exome studies demonstrated the ability to identify causal alleles for a variety of inherited diseases (Bilgüvar et al., 2010; Ng et al., 2010). In rare cases, sequencing of patient samples has suggested specific clinical interventions that have dramatically altered patient outlook. In one early example, exome sequencing of a child with severe inflammatory bowel disease uncovered a mutation in an important regulator of inflammation, X-lined inhibitor of apoptosis (XIAP). Based on the severity of the child’s symptoms as well as the molecular diagnosis, a bone marrow transplant was given to the patient, which subsequently alleviated his symptoms (Worthey et al., 2011). Despite the power of HTS for disease gene discovery, however, exome sequencing currently identifies the genetic defect in only 25% of cases (Yang et al., 2013).
Cancer genome sequencing
Cancer is another important arena where HTS has been applied to great effect. The Cancer Genome Atlas (TCGA) and International Cancer Genome Consortium (ICGC) have performed genome and exome sequencing on thousands of tumor-normal pairs. These studies have described the mutational landscapes for over 20 cancer types, demonstrating that tumors can vary dramatically in both the type and quantity of mutations (Chang et al., 2013; Lawrence et al., 2014). These global descriptions have been integral to the development of background mutation rates that are necessary for the detection of cancer driver genes. For example, replication timing and gene expression were both found to be important covariates when determining if a gene is mutated at a rate higher than expected (Chang et al., 2013). Using these background models, TCGA-led projects discovered several novel cancer drivers, known drivers in new cancer types and commonly disrupted pathways (Lawrence et al., 2014). Moreover, WGS of cancer samples has also identified high-frequency, non-coding mutations, such as activating mutations in the TERT promoter (Huang et al., 2013), a poorly characterized but highly relevant class of somatic variants.
The scale and sensitivity of HTS has also enabled global descriptions of tumor heterogeneity, clonal evolution and the mechanisms underlying drug resistance. By tracking copy number aberrations in primary breast cancer cells using single-cell sequencing techniques, Navin and colleagues demonstrated that copy number rearrangements can occur in bursts, followed by persistent clonal expansion (Navin et al., 2011). Point mutations, in contrast, appear to accumulate more slowly over time, giving rise to more extensive clonal diversity, which may enable the tumor to adapt to diverse selective pressures (Wang et al., 2014). In addition to examining clonal diversity, HTS has also been used to compare primary tumors with relapse lesions, allowing characterization of the effects of chemotherapy as well as the molecular mechanisms underlying resistance to therapy (Van Allen et al., 2014; Ding et al., 2012). Together, these molecular portraits of cancer are forming the foundation of new paradigms for the diagnosis and treatment of cancer.
Limitations of current HTS technologies
It is becoming increasingly clear that while the technologies of today may be capable of providing population-level sequencing to both researchers and clinicians, key limitations remain. From a technological perspective, accuracy and coverage across the genome are still problematic, particularly for GC-rich regions and long homopolymer stretches (Ross et al., 2013). In addition, the short read lengths produced by most current platforms severely limit our ability to accurately characterize large repeat regions, many indels and structural variation, leaving significant portions of the genome opaque or inaccurate (Snyder et al., 2010). The establishment of a gold standard genome, as envisioned by the Genome in a Bottle Consortium (Zook and Salit, 2011) as well as standards for data processing, variant calling and reporting as set out in the CLARITY Challenge (Brownstein et al., 2014), will be valuable for comparing and reporting the accuracy of different platforms and studies. Given the limitations and biases of different platforms, it is also likely that accurate genome sequencing will use a combination of technologies.
In addition to genomes, quantitative analysis of complete transcriptomes, with individual allelic and spliced isoforms, is hindered by short reads as well as the cost and throughput of current long-read technologies. Improvements to current long read technologies, such as Pacific Biosciences and Oxford Nanopore Technologies, as well as the use of “synthetic long read methods” in which longer fragments can be sequenced and assembled from short reads will help overcome these limitations (Tilgner et al., 2015). Although both the research and medical communities are pressing forward with current technologies, these limitations will also continue to drive the innovation of new sequencing platforms (reviewed by Schadt et al., 2010).